Products & Applications
Sequins Metagenomics Core Control Set
Industry leading quantification standards for microbial populations.



Most Comprehensive Metagenomics Control Set
The Sequins™ Metagenomics Core Control Set is a comprehensive, easy-to-use, pre-configured control set with built-in redundancy that ensures integrity of results. It contains synthetic sequences that collectively represent 52 microbial species at varying abundances to emulate the complexity found in a natural microbial community.
Features include: quantitative ladder; representation of domains including bacteria, archaea and eukaryotes; GC% content range (27% to 71%); single tube blend; demonstrated on various samples using Illumina and Oxford Nanopore technologies.
Genus |
No. Species |
Genus |
No. Species |
Genus |
No. Species |
| Aciduliprofundum | 1 | Fusobacterium | 1 | Persephonella | 1 |
| Acinetobacter | 2 | Helicobacter | 1 | Porphyromonas | 1 |
| Anabaena | 1 | Klebsiella | 1 | Roseobacter | 1 |
| Bacillus | 2 | Lactobacillus | 1 | Salmonella | 1 |
| Buchnera | 1 | Legionella | 1 | Shigella | 1 |
| Caldicellulosiruptor | 1 | Leuconostoc | 1 | Staphylococcus | 1 |
| Candida | 1 | Listeria | 2 | Streptococcus | 1 |
| Candidatus Caldiarchaeum | 1 | Magnetococcus | 1 | Synechocystis | 1 |
| Candidatus Carsonella | 1 | Metallosphaera | 1 | Thermotogan | 1 |
| Candidatus Korarchaeum | 1 | Methanococcus | 1 | Thermus | 1 |
| Chlamydia | 1 | Methanopyrus | 1 | Toxoplasma | 1 |
| Chlorobium | 1 | Mycobacterium | 1 | Treponema | 1 |
| Corynebacterium | 1 | Neisseria | 2 | Vibrio | 1 |
| Desulfitobacterium | 1 | Nitrosopumilus | 1 | Vulcanisaeta | 1 |
| Desulfovibrio | 1 | Nostoc | 1 | Wolinella | 1 |
| Ehrlichia | 1 | Oenococcus | 1 | Yersinia | 1 |


Best In-Class Quantification & LOD Determination
Sequins delivers best-in-class quantification and LOD determination, with a highly reproducible molecular ladder, broad dynamic range, and reliable performance across complex sample types and sequencing platforms. Its strong quantitative accuracy, clear assay sensitivity readout, and negligible sample interference make it a robust in-sample ground truth for metagenomic workflows.
This figure provides an overview of the analytical performance of the Sequins Metagenomics Core Control Set, showing how it supports sensitive detection, accurate quantification, and reliable use across different sample backgrounds and sequencing platforms. Collectively, the data demonstrate the value of Sequins as a robust in-sample reference for quantitative metagenomics and overall assay performance assessment.
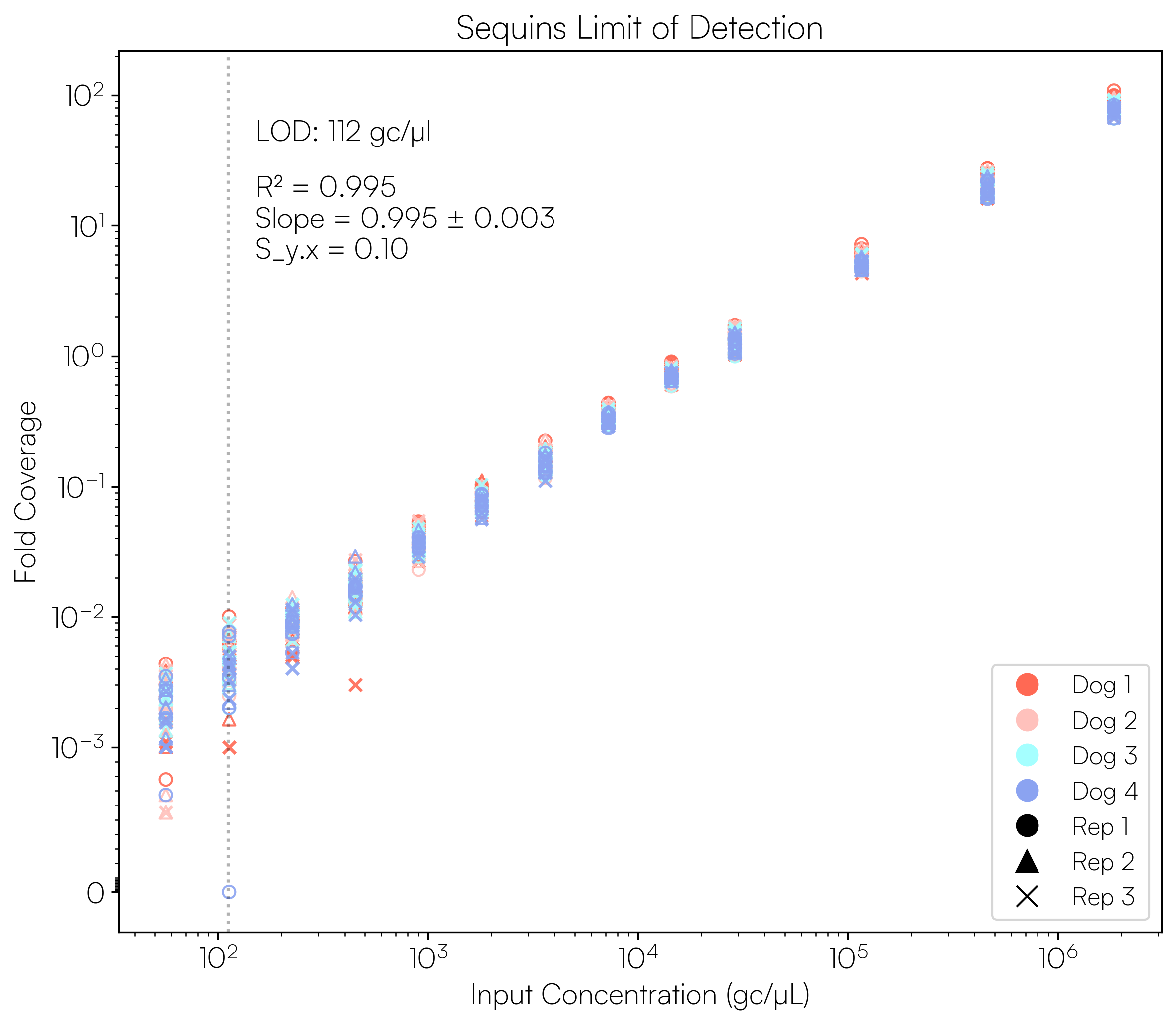
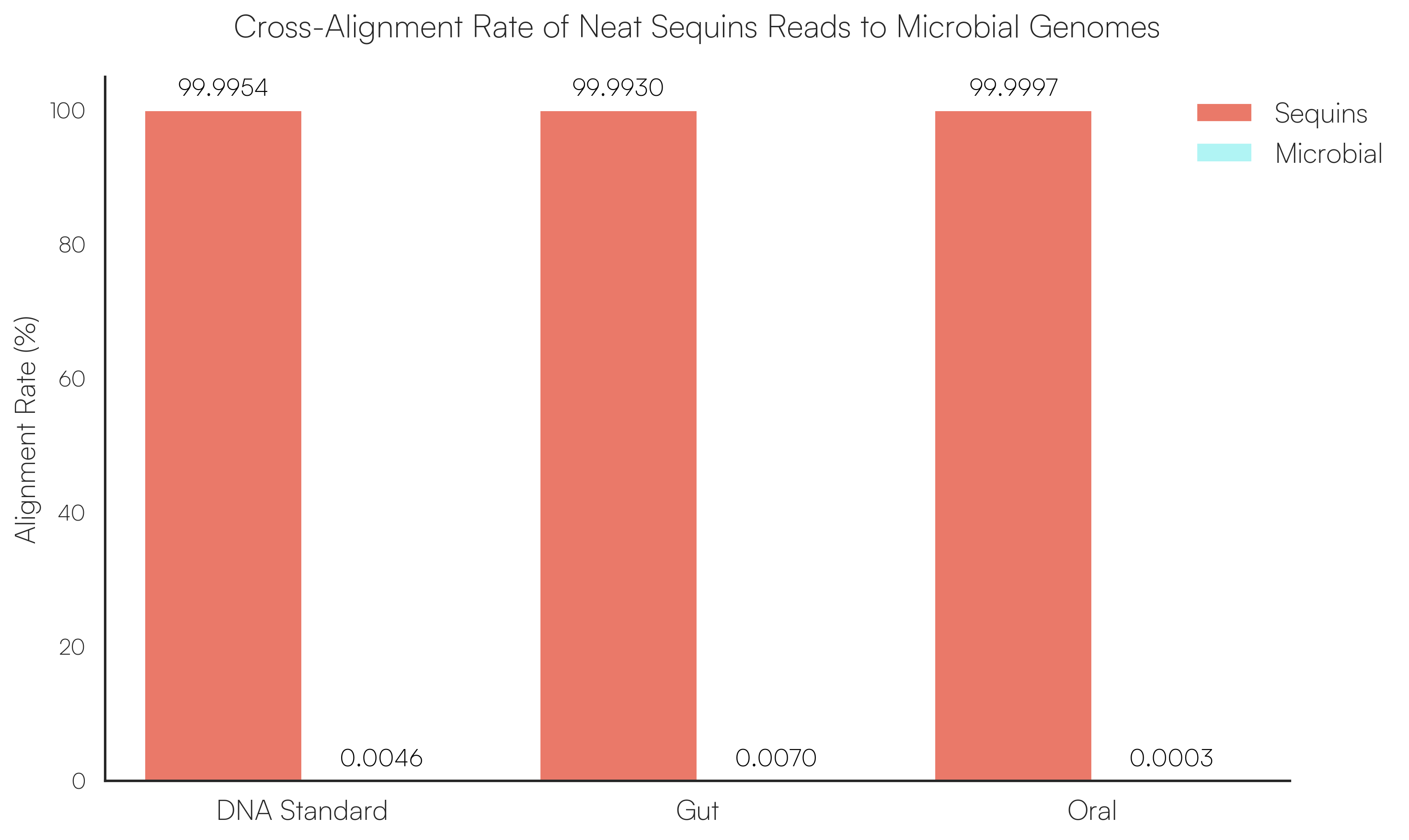


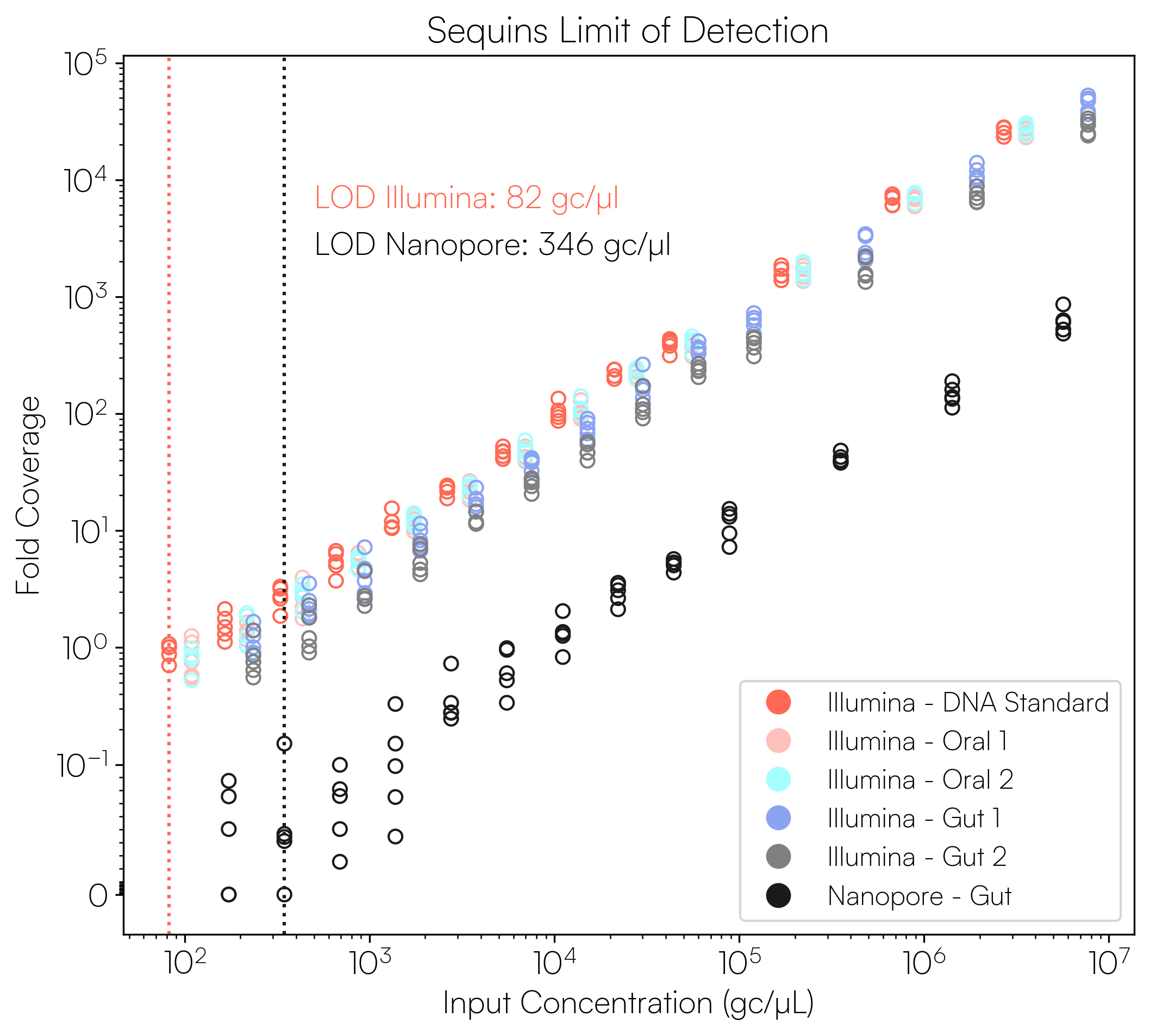
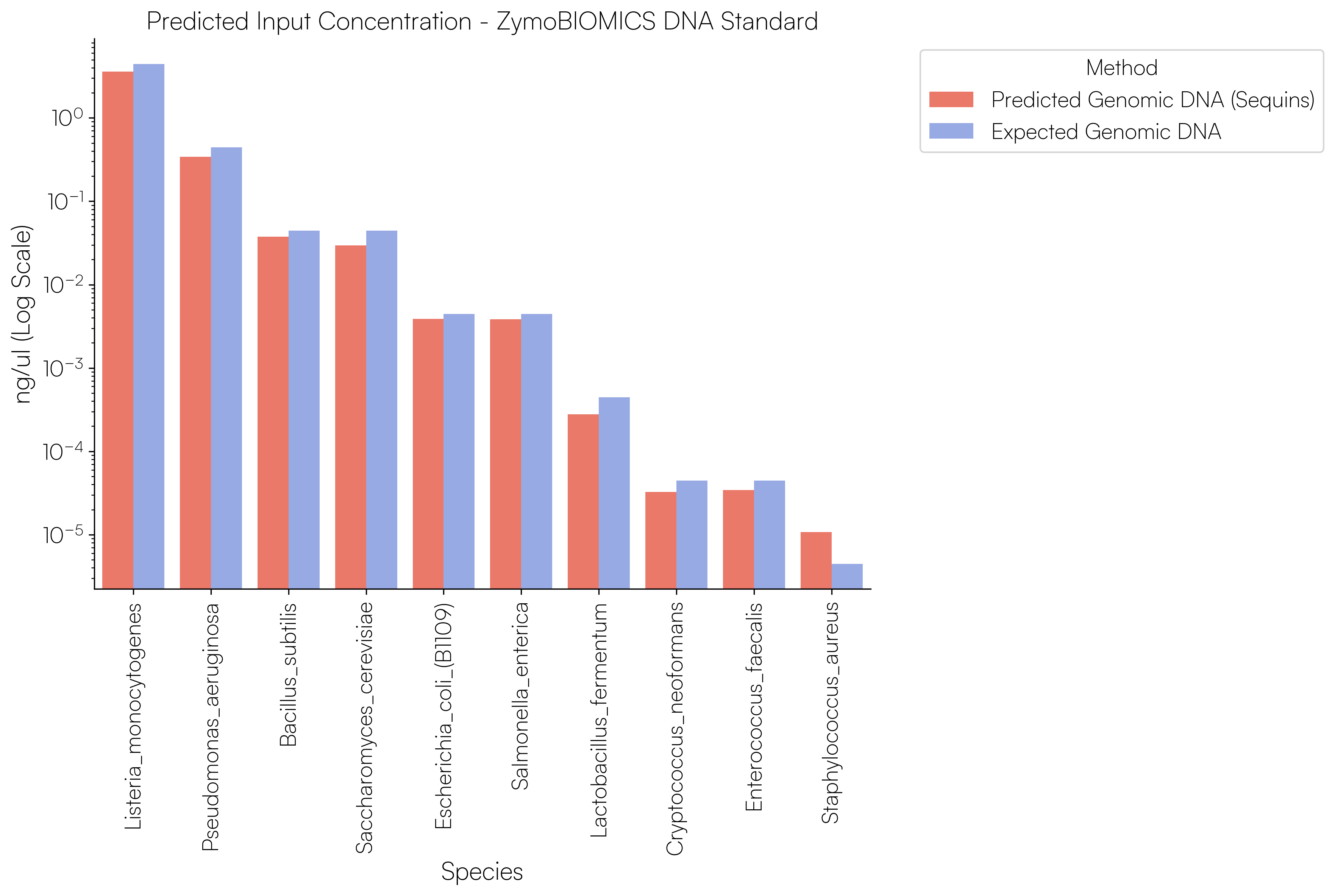


Sequins Metagenomics Core Control Set performance. (A) Metagenomics Core Control Set replicates in dog faecal matrix background demonstrating an accurate and reproducible ladder in complex samples. (B) Cross-alignment rate of Sequins reads to microbial genomes, demonstrating that Sequins does not interfere with samples – less than 0.01% of reads cross-align to mock community genomes. (C) Sequins ladder on various backgrounds generated using Illumina and Oxford Nanopore Technologies (ONT) sequencing. This demonstrates that Sequins is compatible with short- and long-read sequencing technologies and can be used to determine assay limit of detection (LOD). (D) Predicted input concentration estimates (predicted using Sequins v. expected based on ZymoBIOMICS DNA Standard) demonstrating that the Sequins molecular ladder enables reliable microbial abundance quantification. Internal data.