Technology
Mirroring Technology & Innovative Design
Maximize data insights across a range of applications, collaborations and locations with simple to use sequencing spike-in controls.



Sequins™ are synthetic nucleic acid sequencing spike-in controls that directly mirror naturally occurring sequences
Because Sequins retain the same nucleotide composition as the natural sequence, they enable accurate representation of genomic complexity without compromising integrity of the sample and results. Sequins perform equivalently throughout sequencing workflows, providing a true measure of control.

Sequins Design Concept
Sequins are rationally designed synthetic reference molecules developed to mirror genomic targets of interest. Depending on the application, Sequins can be formulated as germline controls, somatic controls, or molecular ladders.

Germline controls
For germline applications, Sequins are designed as matched pairs, with one molecule representing the wild-type allele and the other the variant allele. These molecules are typically blended in a 1:1 ratio, resulting in an expected 50% variant allele frequency, consistent with a heterozygous state.
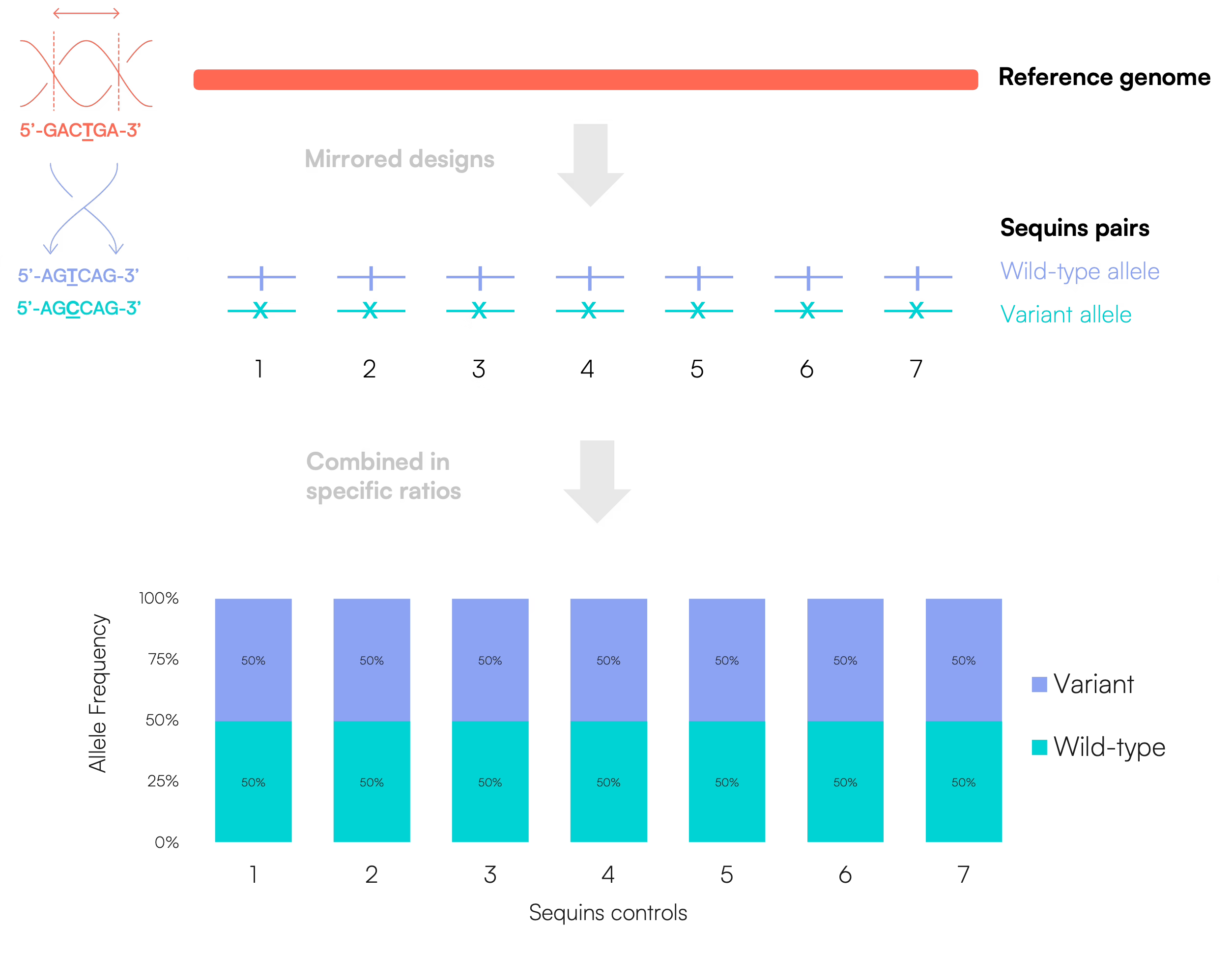

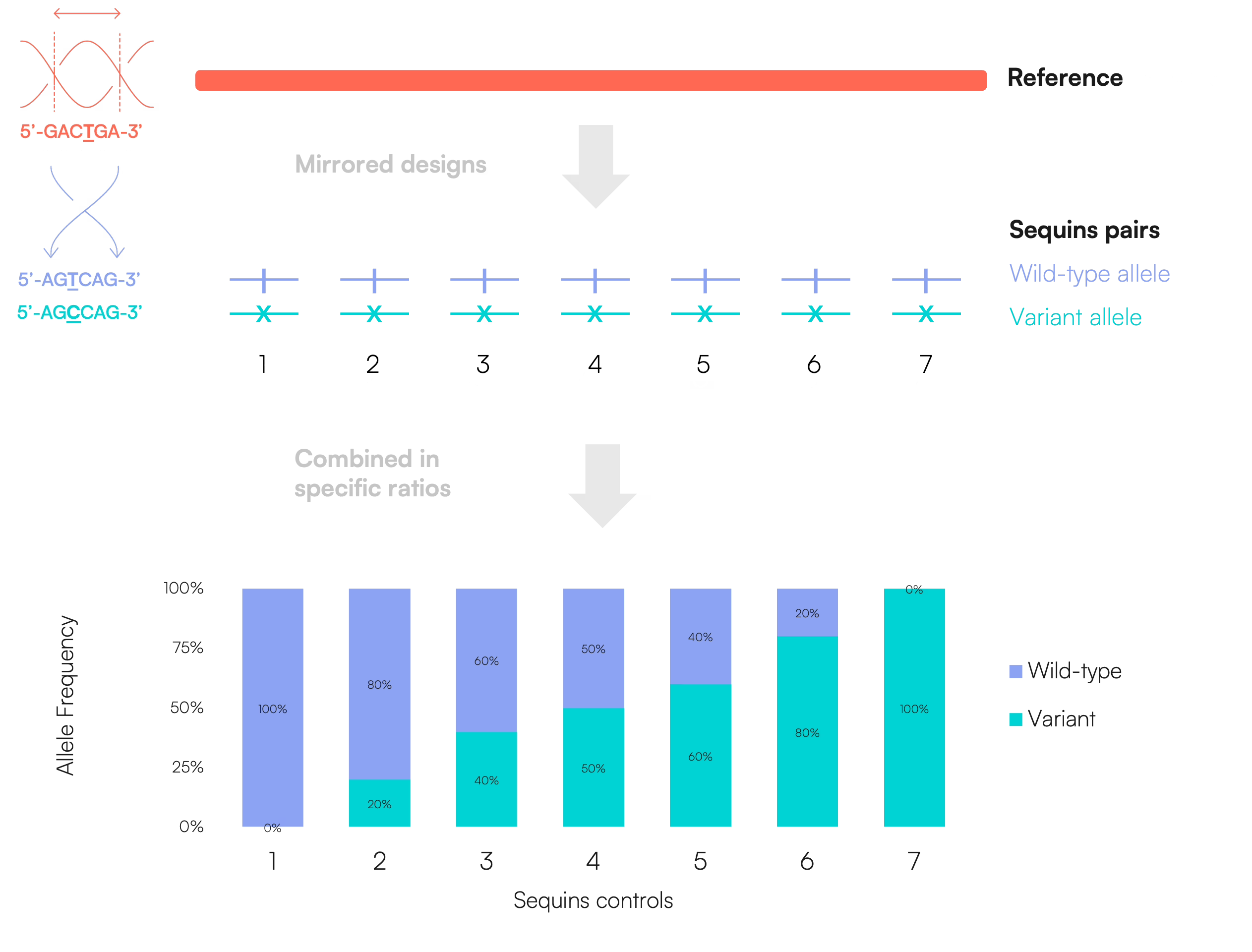
Somatic controls
For somatic applications, Sequins are also designed as matched wild-type and variant pairs, but can be blended at defined ratios to generate the desired variant allele frequencies. This enables the creation of controls representing low-, medium-, or high-frequency variants for assay development, validation, and performance assessment.

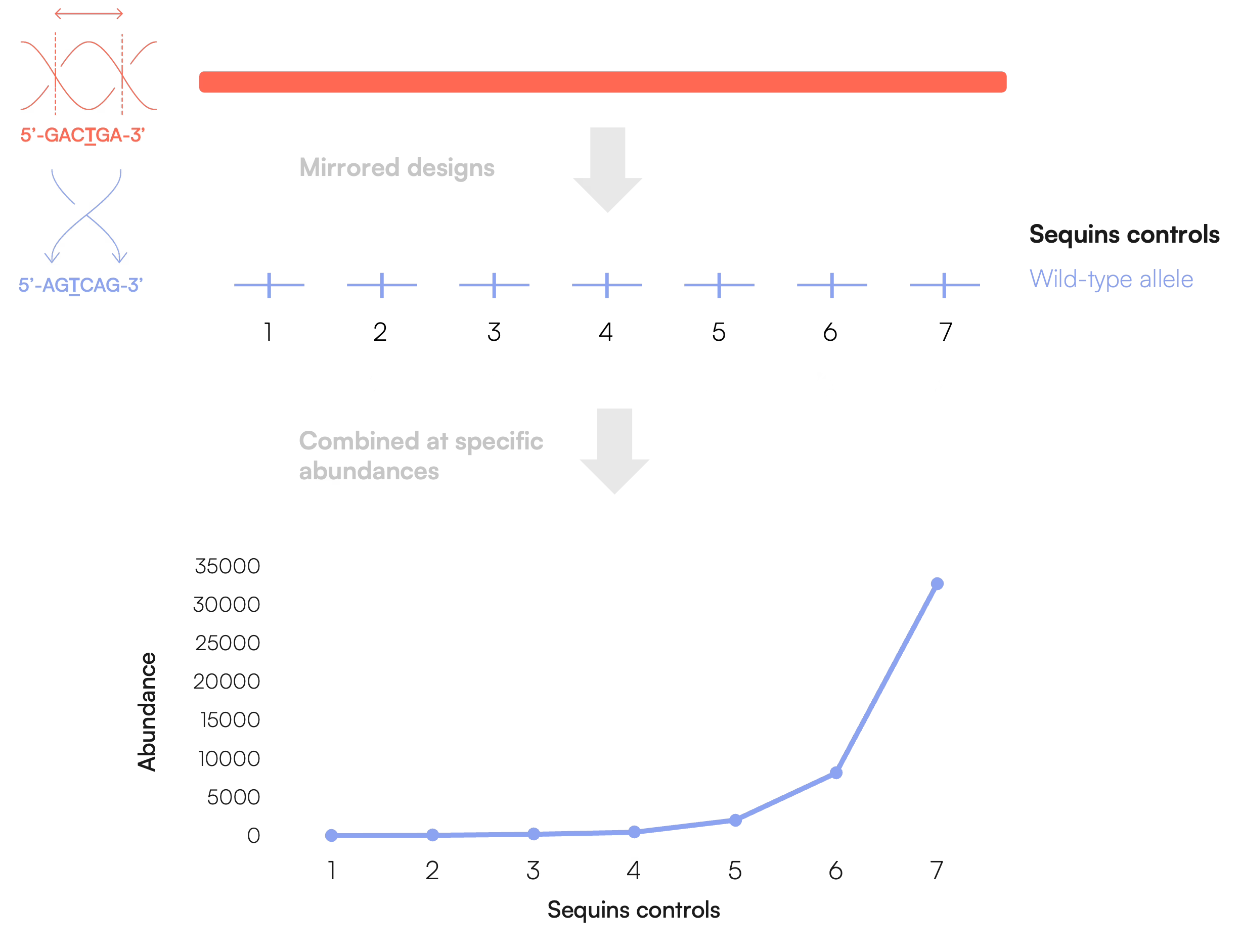
Molecular ladders
Molecular ladders are constructed using a set of individually designed mirrored Sequins molecules each representing a distinct target region and is designed to be distinguishable from both the native sample and from other Sequins molecules within the ladder.
Sequins molecules are blended at defined concentration or abundance levels to create a series of quantitative reference points. Importantly, each point on the ladder is typically represented by multiple distinct Sequins molecules rather than a single molecule alone. This helps improve robustness, reduces dependence on the behaviour of any one sequence, and minimises sequence-specific bias.
The resulting molecular ladder functions as a standard curve for quantitative analysis, supporting calibration, assessment of linearity and dynamic range, and more confident comparison of sample measurements across an assay or workflow.